<a href="http://www.shutterstock.com/cat.mhtml?lang=en&search_source=search_form&version=llv1&anyorall=all&safesearch=1&searchterm=bengal+tiger&search_group=#id=62798404&src=_EYKoNheKT4-e1Skr16xjQ-1-39">Sergey Uryadnikov</a>/Shutterstock
Update: Kathmandu-based reporter Kashish Das Shrestha was also along on this reporting venture, and has published his story on the Tiger Genome Project on his website, Sustainable Nepal.
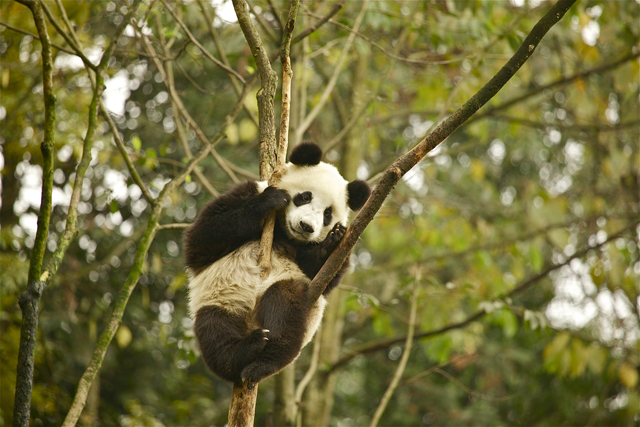
Bengal tigers can be elusive. They’re classified as an endangered species, they’re mostly nocturnal, and if they had their way, they wouldn’t see many humans, either. Native to Southeast Asia, there are only an estimated 1,850 left in the wild. That makes counting them somewhat difficult—but researchers in Nepal have developed a system that they think will make it easier to figure out how many tigers live there. They’re pulling genetic data out of their poop.
Founded in 2011, the Nepal Tiger Genome Project has collected more than a thousand scat samples from the southern part of the country known as the Terai Arc landscape, one of the last remaining tiger habitats on the earth. Not to get too graphic, but when tigers do their doo, it sloughs off some of their cells on the way out, from which scientists can extract DNA. The DNA allows the researchers to study and catalog the genetic material and to create a database of all the country’s tigers.
Dibesh Karmacharya is the executive director of the project, which he runs through his biotechnology company, Intrepid Nepal, and the Center for Molecular Dynamics-Nepal, a research organization that he also directs. Karmacharya returned to Nepal after 14 years in the US working in biotech, and started the lab to focus mainly on molecular diagnostics for human diseases. The lab’s work on the Tiger Genome Project brings together two things Karmacharya loves—animals and genetics. “I wanted to be a wildlife photographer,” he told me in his office in Kathmandu last week. “I could never get a job doing wildlife in the US. I ended up getting a job in genetics, because that was my skill.”

The wildlife genetics work started when the World Wildlife Fund asked the lab to help track snow leopards, a threatened species native to central Asia. After seeing the success of the snow leopard work, Karmacharya and several researchers from the US—Marcella Kelly, an associate professor in the Department of Fish and Wildlife Conservation at Virginia Tech, and Lisette Waits, a professor in the Department of Fish and Wildlife at the University of Idaho—proposed the Tiger Genome Project and secured a $270,000 grant from the US Agency for International Development (USAID) to fund the initial work. (Full disclosure: I was in Nepal to help with a USAID-sponsored environmental reporting workshop.)
To gather the samples, the Tiger Genome Project sent surveyors—armed with specimen vials and field surveys for logging the GPS location, type of forest cover, and condition of the scat— into four national parks and the wildlife corridors that tigers are thought to use to pass between parks. Project leaders hoped to collect 700 samples, but the crew turned up 1,200 over the course of more than two months. “We collected a lot more shit than we thought we would,” Karmacharya joked.
Back in the lab, Karmacharya and his research team extract and amplify the DNA from the scat samples, which is actually pretty difficult. The scat can be in various stages of decay, and may also contain segments of DNA from whatever the tiger ate for dinner. But from the extracted DNA, they can usually determine the species and sex of the animal that left the scat behind.
Karmacharya pulls one of the vials of scat from a closet and hands it to Manisha Bista, the project coordinator. She scans the bar code on the vial, and a computer searches their database to pull up the information cataloged on the producer of this scat. Scat #485, found in Chitwan National Park, was male. On another screen a Google map shows where the scat was found, and that they have identified multiple samples from the same animal. The lab expects to finish processing all the samples by the end of May; so far, they have identified 50 distinct tigers from their scat. The 2012 survey of Nepal found a total of 176 tigers throughout the country.

This isn’t the first research project tracking the DNA of animals from their droppings; this type of genetic evaluation started back in the early 1990s with research on bears, and has also been used on tigers in India. But it’s still a relatively rare method—there are only about 20 labs in the world doing this work, according to Waits—and this is one of the largest and most comprehensive surveys of its type to date.
A more traditional method of tallying tigers uses heat- and motion-detecting cameras posted throughout the known habitats. This, Kelly says, is probably still the most cost-effective and efficient method. But using camera traps and DNA collection together gives researchers new insight into the animals, allowing them to evaluate the genetic diversity and health of population, the range that the tigers travel, and how genes flow between populations. It can help conservation specialists carve out protected areas based documented evidence of the tigers’ range and preferred habitat.
The Tiger Genome Project is also useful in prosecuting poachers, who sell pelts and bones for trophies and ceremonial use. If part of a tiger is confiscated from poachers or wildlife traders, researchers can use its DNA to see if it’s been previously cataloged in their system. Last month, Nepali law enforcement asked Karmacharya to ID two skin samples from a seizure; the DNA didn’t match any of the tigers that they’ve cataloged, but it demonstrated the potential of the technology.
The biggest benefit, researchers say, is the potential to study animals for which it is notoriously difficult to get genetic material. “One of the main benefits is you don’t have to capture or handle the animals, which puts animals at risk and people at risk—particularly when you’re dealing with large carnivores,” says Waits, who in addition to working on the Tiger Genome Project, has worked with DNA from scat samples for all kinds of animals, from brown bears to large cats. “For species that are rare or difficult to capture, they still have to poop.”